Scope: On methodology for assessing potential presence of Treatment Effect Heterogeneity [HTE] in randomized and observational data. (This SIG was formally known as the "Subgroup" SIG, a less accurate name).
We are: Arkan Abadi, Björn Bornkamp, Daniel Bratton, Mathias Cardner, Eliana Garcia Cossio, Paulo Eusebi, Frauke Hennig, Jared Huling, Thomas Jemielita, Serene Jiang, Jeremiah Jones, Runjia Li, Ilya Lipkovich, Zihuan Liu, Karl Koechert, Nicole Kraemer, Henrik Loft, Brian Millen, Esha Mohammed, Heiko Goette, Lukas Lasoto, Bohdana Ratitch, Marie-Karelle Riviere, Barbara Rosettani, Kostas Sechidis, David Svensson, Julien Tanniou, Marius Thomas, Lou Whitehead, Huang Xin, Xu Yuejia, Pingye Zhang. [Updated June 2025]
The SIG lead is David Svensson (david.j.svensson@astrazeneca.com).
_______________________________________________________________________________________________________
The assessment of potential differential treatment effect arise in many ways in pharmaceutical development, sometimes more as a regulatory-risk problem (e.g., to demonstrate consistency of treatment effect across a pre-defined set of subgroups) and sometimes more as a sponsor's risk (selection of a subgroup to be put forward as the basis of another trial). A purely exploratory assessment is also commonly considered, as an attempt to try to 'learn a lesson' from a trial which perhaps failed to meet it primary (overall) objectives.
There are well-known inherent statistical difficulties with all the above; with consistency, due to limited data in the subgroups, there is a high risk of false positives (random highs) as well as a low power to detect true differential effects (since trials are seldomly sized for it). It is worth mentioning that consistency assessment is routinely performed without strong error rate control (since it otherwise would require dramatically larger sample sizes). In the subgroup selection setting, it is of key importance to provide an honest estimate discounted for the number of subgroups inspected, in order to not overstate the real effect.
It should be stressed that the assessment of Treatment Effect Heterogeneity [HTE] goes beyond merely inspecting subgroups pre-specified in a protocol. Many recent techniques are data-driven causal inference approaches with a Machine Learning [ML] flavour, and here some confusion sometimes can arise: the assessment of a potential presence of treatment effect heterogeneity should not be phrased as a standard ML problem (see e.g., our 2024 Tutorial paper in Statistics In Medicine: https://lnkd.in/gJafkXCn co-written by SIG members). For one thing, the ground truth - the Individual Treatment Effects [ITE] - are fundamentally unobservable, therefore ruling out any standard application of Supervised Learning ML. Instead, the problem has to be phrased within a potential outcome framework, where ML can enter as modelling components, where the key aspect is not so much the choice of ML methods themselves (such as e.g., RandomForest vs Boosting) but primarily on the choice and impact of the architecture regarding how the models are combined for rendering estimates of CATE (the Conditional Average Treatment Effect). Many inherent research questions are only partially resolved, e.g., which architecture results in best performance (e.g., regarding regularization biases, ability to correctly rank predictive biomarkers, enable omnibus tests of HTE).
The area is progressing fast and (in fact) other industries are interested too in estimating causal effects, with interesting methods coming from their direction. Therefore, it makes sense to broadly keep an eye on methodological developments. There is still a long list of more or less unresolved theoretical problems; also, many practical considerations merit a lot of attention, e.g., well described in the following paper co-written by several SIG members (with other external experts involved): https://lnkd.in/gfPzpKxJ. This paper ought to interest many.
The SIG should be considered as a network of researchers in this area with primary aim to promote knowledge sharing, collaborations and awareness across the industry. Currently, several papers are in the making by self-organising subgroups of SIG members, happening above our regular all-hands meetings (where we tend to discuss recent papers, sometimes presented by guest researchers). The SIG often co-authored presentations for various conferences, examples inserted below. The latest example is the PSI conference in London where we have a session (10th June) focusing on the notoriously difficult problem of assessing consistency of treatment effects across subgroups as well as looking at group sequential design options for a setting where there is a subgroup involved in the primary testing. (Speakers: Björn and Marie, David moderator).
A recent collaboration among many SA-SIG members and other experts involved resulted in a practical process-oriented guideline for a structured approach to of HTE assessment, including preparation of data, cross-functional communication with stakeholders, and interpretation of findings. K. Sechidis, S. Sun, Y Chen, J. Lu, C. Zhang, M. Baillie, D. Ohlssen, M. Vandemeulebroecke, R.
Hemmings, S. Ruberg, B. Bornkamp. "WATCH: A Workflow to Assess Treatment Effect Heterogeneity in Drug Development for Clinical Trial Sponsors" (https://onlinelibrary.wiley.com/doi/10.1002/pst.2463?af=R)
Another recent SA-SIG collaboration resulted in a tutorial-style overview of recent trends in the literature, covering how the methodology evolved since the previous tutorial paper by Lipkovich et. al 2016. This paper focus on causal inference based modelling for estimating CATE (Conditional Average Treatment Effect) and related Treatment Regimes, as well as overviewing such aspects as post-selection inference (e.g., so called nGATES inference, omnibus tests of heterogeneity, etc). I. Lipkovich, D. Svensson, B. Ratitch, A. Dmitrienko, Modern approaches for evaluating treatment effect heterogeneity from clinical trials and observational data, Statistics in Medicine, 2024.
Another publication compares the various methods for assessing treatment effect heterogeneity, but also evaluates their performance in simulation scenarios that mimic real clinical trials. Furthermore it introduces an R package (benchtm), which can simulate synthetic biomarker distributions based on actual clinical trial data and create interpretable scenarios to benchmark methods for identifying and estimating treatment effect heterogeneity. S. Sun, K. Sechidis, Y. Chen, J. Lu, C. Ma, A. Mirshani, D. Ohlssen, M. Vandemeulebroecke, B. Bornkamp, Biometrical Journal, 2024.
Furthermore, the following paper was submitted in 2025, co-written under the umbrella of the SIG: https://arxiv.org/abs/2505.01145. The paper explores SHAP application for identifying predictive biomarkers through CATE models. In particular, inherent challenges associated with the SHAP concept in multi-stage CATE strategies is discussed and a surrogate estimation approach is proposed that is agnostic to the choice of CATE strategy, effectively reducing computational burdens in high-dimensional data.
BELOW: Further material (including some presentations) which gives a flavour of the scope of the SIG.
SIG_TreatEffHet_posterBasel2024
Lipkovich_bbs_2024_final
DS_etal_PSI2024
2024_06_PSI_Sechidis
PSI2023_14.06_Yuejia_Xu
psi 2023_PauloEusebi_etal_Data-driven subgroup identification in clinical trials
Subgroup_Challenge_BB (PSI 2022)
PSItalk2021_DS_etAl_Consistency

Examples of our Meetings (not complete list):
- 2018 APRIL: Update re progress of White Paper, remaining work for 2018. Ideas include further work on Bayesian shrinkage, Model averaging, Simulations under NULL when prognostic factors are present, SEAMOS development for non-linear models and Bootstrap Bias reduction.
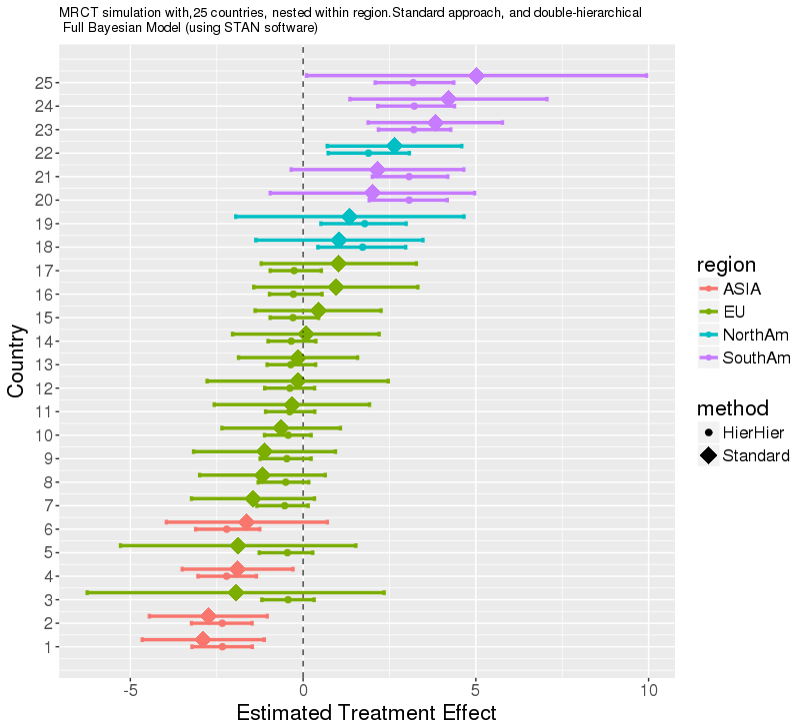
- 2018 JUNE: David PSI presentation on some aspects of Shrinkage, multi-level hierarchical models, and model averaging. Key aspect: many variations exist, and some unknowns regarding performance.
- 2018 JULY: Simulation of RCT discussed with underlying prognostic predictive continouos variables - dichotomized into subgroup factors - some preliminary illustrations of methods listed under APRIL.
- 2018 SEPT: Further simulations on the performance of BIC model averaging.
- 2018 NOV: Visualisation using novel R package SubrPlots (e.g., UpSet graph), Amy Spencer presenting further work on SEAMOS (modification to increase power). Updates on PSI 2019 Subgroup Section
- 2019 JAN: Discussing content for SIG subgroup session at PSI, presentation by David Svensson on SEAMOS (some simulation results in a non-linear case with prognostic factors).
- 2019 MARCH: Review of novel R CRAN package 'subtee', containing functionality for Bootstrap-based bias-reduction and BIC-Model-Averaging Shrinkage. Some simulation examples were discussed, and it was decided that more such (across broader assumptions) will be the topic for another SIG meeting. Issues regarding hierarchical Bayesian shrinkage when subgroups are overlapping was also discussed, with particular attention to a paper by Varadhan and Wang.
- EFSPI WORKSHOP 20th MARCH 2019 on Recent Developments on Subgroups and Biomarkers (Gothenburg) saw contributions from the SIG (presentations by G. Rosenkranz, and I.Lipkovich & D.Svensson) with the focus on HTE/Individual Treatment Effects, but also general aspects of consistency across subgroups was discussed, including some regulatory contribution.
- MAY 2019: Discussion of resampling principles for creating NULL data (idea: generating a large number of replicates with marginal distribution and correlations as in observed data except that treatment hetereogeneity is taken out); discussings also regarding possibities of a new points-to-consider document for exploratory analyses ('reality check' aspects to consider when assessing a given datamining/subgroup detection analysis).
- JUNE (PSI LONDON 2019): SIG members presented an overview and some recommendations regarding subgroup detection (Necdet Gunsoy, Ilya Lipkovich & David Svensson) at the SIG session (chaired by Aaron Dane).
- AUGUST 2019: Further discussions and planning of compiling a points to consider document that would help upfront when planning/conducting/reviewing an exploratory subgroup analysis (the latter to be interpreted in a broad sense, including data-driven approaches). Discussions regarding the upcoming PSI webex and PSI Subgroup session for Barcelona 2020.
- SEPTEMBER 2019: discussing best practices when performing ML SubId, initiating a long term SIG work in this direction. Common difficulties, common likely errors and how to avoid it.
- NOVEMBER 2019: further work on the above, planning a joint PSI talk in this direction.
- DECEMBER 2019: Further work on the joint presentation & possible SIG white paper.
- JANUARY 2020: on simulations for illustrating issues to avoid/overcome in SubId in the practice
- MARCH 2020: On formal Type 1 Error control in ML - novel ideas
- MAY 2020: Presentation and group discussions of a novel idea for SubId in Dose Response (paper by Marius Thomas, Bjoen Bornkamp, Heidi Seibold).
- JULY 2020: overview of recent ideas in the ITR area; optimizing Treatment for a given patient.
- OCTOBER 2020: ITR analyses in Multi-Treatment trials.
- NOVEMBER 2020: Knock-Offs Variable Importance (Kostas Sechidis)
- JANUARY 2021: Virtual Twins implementation aspects (shared surface, XGboost) and Causal Forest (David Svensson)
- MARCH 2021: Assessment Evaluation of different analytic strategies for estimating optimal treatment regimens for time-to-event outcomes in observational data (Ilya Lipkovich)
- 6th JUNE 2021: Presentation by Oliver Keene on Subgroup Dynamic Borrowing.
- 22th JUNE 2021: SIG SESSION at the PSI Conference (speakers: D. Svensson, D. Lawrance)
- 14th OCTOBER 2021: Discussing simulations regarding SEAMOS vs classical methods, beyond linear models (M. O'Kelly, D. Svensson)
- 17th NOVEMBER 2021: PSI SIG WEBINAR: Modern approaches to subgroup identification. (speakers: Svensson, Lipkovich, Bornkamp, Sechidis, Eusebi).
- 2nd DEC 2021: Discussing role and design of simulations for evaluation of methodologies.
- February 2022: group discussions of some recent papers (Modified covariate methods, value based for survival data)
- March: discussing BART; Bayesian Causal Forest (D. Svensson)
- May: Yuejia presented her Ph.D Thesis re optimal ITR in multi-arm settings
- June, 13: PSI Gothenburg SIG session; speakers David Svensson, Björn Bornkamp and Stefan Franzen.
- September, 26: SIG meeting covering PSI 2022, and a potential 2023 session, and ideas for collaborations.
- October 2022: Daniel Bratton - presentation of new ideas re Shrinkage without exchangeability assumptions
- November 2022: Nicole Kraemer - presentation of a resampling based approach for Consistency assessment
- December 16th 2022: Biomarker SIG Co-Leads visiting Subgroup SIG - connecting/sharing ideas
- January 2023: David Svensson presenting simulation discovery regarding Causal Forest - joint work with Lipkovich and Ratitch.
- June 2023: PSI SIG session (speakers: Paulo Eusebi, Nicole Kraemer, Yuejia Xu; all co-work within the SIG)
- July 5th 2023: David S presenting his impressions from the paper "Comparing algorithms for characterising treatment effect heterogeneity" (co-written by SIG members Sun, Sechidis, Bornkamp), group discussions.
- Sept 2023 Mathias Cardner presenting his paper 'Analysis of serum proteomics data identifies a quantitative association between beta-defensin 2 at baseline and clinical response to IL-17 blockade in psoriatic arthritis'
- October 2023: Sophie Sun presenting benchmarking paper comparing a multitude of approaches for discovering predictive biomarkers.
- Dec 2023: Nathalie Weidinger presenting benchmarking re Knockoffs, PPlasso, Shapley Values, Variable Importance, VSURF, SIDES, Virtual Twins, etc.
- Jan 2024: Andrea C presenting simulation benchmarking results concerning subgroup identification.
- March 2024: Zihuan Lie. Neural Networks for Causal inference
- April 2024: Introducing Evaluation of Individualized Treatment Rules: David Svensson.
- May 2024: Evaluation of Individualized Treatment Rules Part 1: Imai Kosuke (Harward).
- June 2024: SIG Session at Amsterdam PSI Conference
- Sept 2024: Evaluation of Individualized Treatment Rules Part 2: Imai Kosuke (Harward).
- October 2024: Stability of Subgroup Identification (Thomas Jemielita)
- Nov 2024: Kostas Sechidis: The WATCH Workflow (SIG paper submitted), discussing recommended ways of approaching the assessment of HTE, in a structured fashion that aims to maximize clarity of results and minimize adhoc data dredging. In particular potential usages of gate-keeper omnibus testing was brought up. (More about this topic in the future).
- January 2025: Vik Shirvaikar (Oxford University) presenting work in progress "Targeting relative risk heterogeneity with causal forests".
- February 2025:David Svensson: presenting selected simulation results regarding performance of CATE approaches from the 2024 Tutorial paper (mentioned above). Approaches considered are meta learners T, S, X, R, Causal Forest (and its Bayesian cousin BCF), Modified Loss Function approaches A and W learning (augmented, non-augmented).
- March 2025: Marie Karelle Riviere, Group Sequential Design options & simulation benchmarking for the case with co-primary (with a BM+ subgroup and full population)
- April 2025: Wilmar Igl, On a novel Shrinkage approach for estimating effect in outlier subgroup (related to consistency assessment)
- May 2025: Bjoern Bornkamp: Benchmarking of various Hierarchical Bayesian Shrinkage approaches (related to consistency assessment).
- June 2025: Eliana Garcia Cossio/ Frauke Hennig / Lukas Lasoto/ Karl Koechert: a proposed workflow for assessing HTE via CATE and SHAP importance (based on CausalForest, R-learning, gatekeeper testing)
- June 2025: PSI SIG session on HTE.